Authors Colin A. Smith csmith@scripps.edu, Ralf Tautenhahn rtautenh@gmail.com, Steffen Neumann sneumann@ipb-halle.de, Paul Benton hpaul.benton08@imperial.ac.uk and Christopher Conley cjconley@ucdavis.edu
Galaxy integration ABiMS TEAM - SU/CNRS - Station biologique de Roscoff and Yann Guitton - LABERCA Part of Workflow4Metabolomics.org [W4M]
Contact support@workflow4metabolomics.org for any questions or concerns about the Galaxy implementation of this tool.
xcms process history
Description
This tool provide a HTML summary which summarizes your analysis using the [W4M] XCMS and CAMERA tools
Workflow position
Upstream tools
| Name | Output file | Format |
|---|---|---|
| xcms.findChromPeaks | xset.RData | rdata.xcms.findchrompeaks |
| xcms.groupChromPeaks | *.groupChromPeaks.RData | rdata.xcms.group |
| xcms.adjustRtime | *.adjustRtime.RData | rdata.xcms.retcor |
| xcms.fillChromPeaks | *.fillChromPeaks.RData | rdata.xcms.fillpeaks |
| CAMERA.annotate | *.annotate.*.RData | rdata.camera.``*`` |
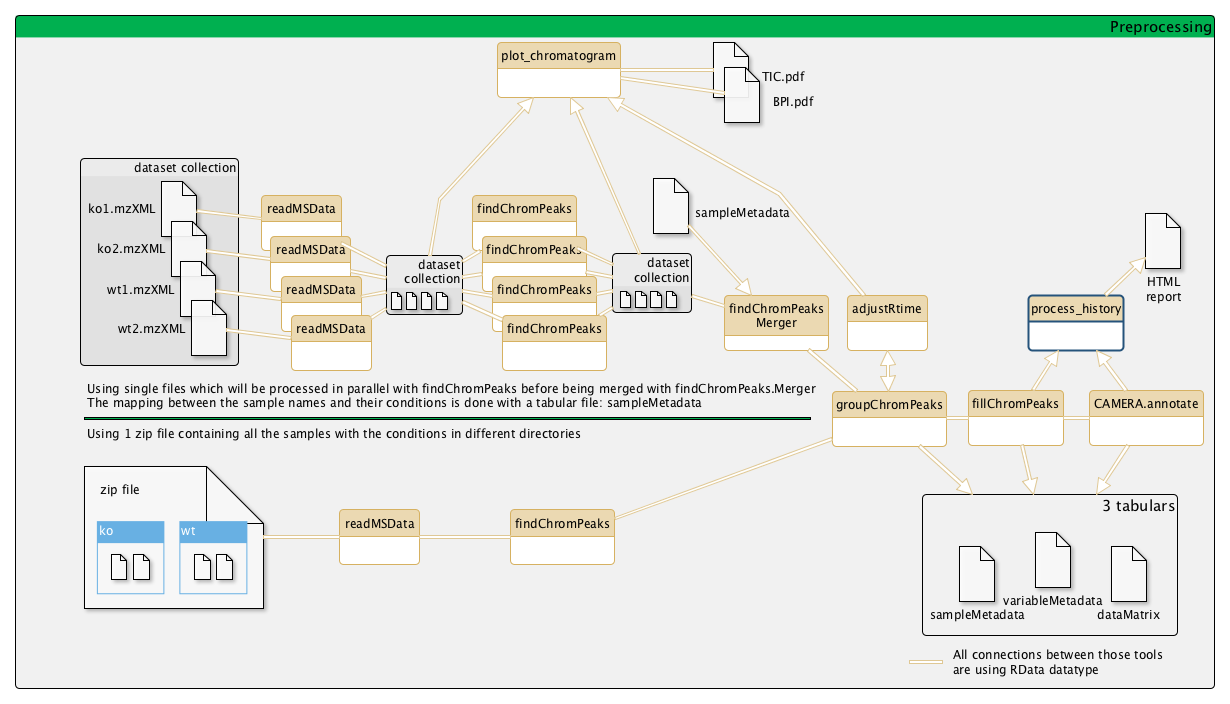
For details and explanations concerning all the parameters and workflow of xcms package, see its manual and this example
Changelog/News
Version 3.12.0+galaxy* - 03/03/2020
- UPGRADE: upgrade the xcms version from 3.6.1 to 3.12.0 (see XCMS news)
Version 3.6.1+galaxy2 - 23/09/2020
- BUGFIX: "Error: object 'sampleNamesList' not found" in case of .RData input from CAMERA
Version 3.6.1+galaxy1 - 05/04/2020
- UPGRADE: upgrade the CAMERA version from 1.38.0 to 1.42.0
- BUGFIX: "Error in chromPeaks(from)[, "is_filled"] : subscript out of bounds"
Version 3.6.1+galaxy* - 03/09/2019
- UPGRADE: upgrade the xcms version from 3.4.4 to 3.6.1 (see XCMS news)
Version 3.4.4.0 - 08/02/2019
- UPGRADE: upgrade the xcms version from 3.0.0 to 3.4.4 (see XCMS news)
Version 3.0.0.0 - 14/02/2018
- UPGRADE: upgrade the xcms version from 1.46.0 to 3.0.0. So refactoring of a lot of underlying codes and methods
- IMPROVEMENT: the tool now shows all the parameters and not only those which were set.
Version 1.0.4 - 13/02/2018
- UPGRADE: upgrate the CAMERA version from 1.26.0 to 1.32.0
Version 1.0.3 - 03/02/2017
- IMPROVEMENT: xcms.summary can deal with merged individual data
Version 1.0.2 - 06/07/2016
- UPGRADE: upgrate the xcms version from 1.44.0 to 1.46.0
Version 1.0.1 - 04/04/2016
- TEST: refactoring to pass planemo test using conda dependencies
Version 1.0.0 - 10/02/2016
- NEW: Create a summary of XCMS analysis