
What it does
FROGS Affiliations stat computes several metrics and generates a HTML file describing OTUs based on their taxonomies and the quality of the affiliations.
Input/output
Input
Abundance file:
The abundance and affiliation of each OTUs in each sample (format BIOM). This file can be produced by FROGS Affiliation OTU.
The FROGS's tools working on clusters and others metagenomic workflows produce files in BIOM format.
Output
Summary file (summary.html):
OTUs taxonomies and affiliations metrics (format HTML):
-Taxonomy distribution: display the distribution of each taxon and the rarefaction for each taxonomic rank and for each sample
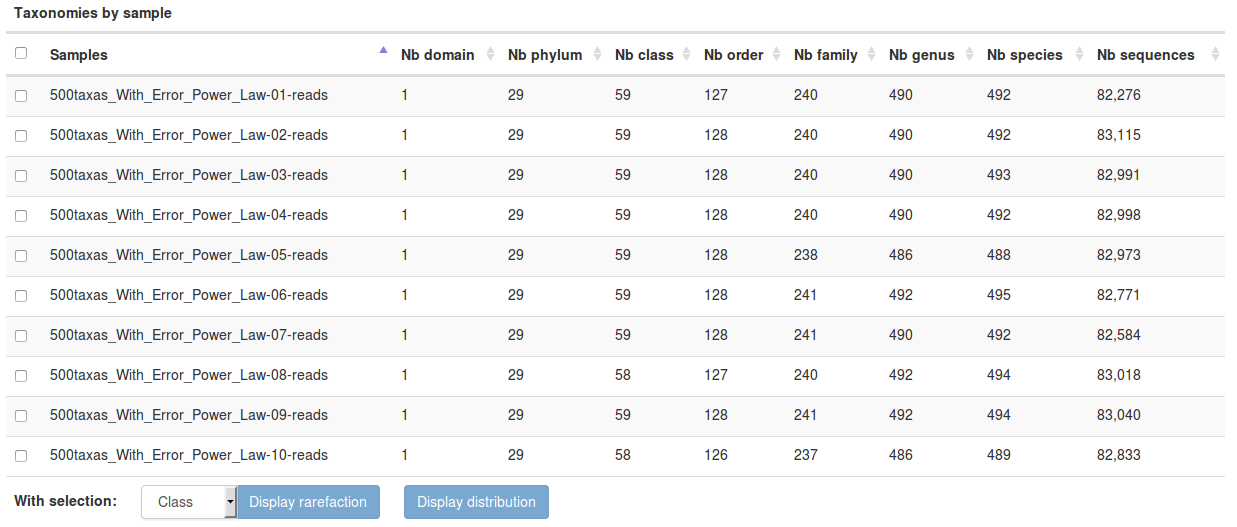
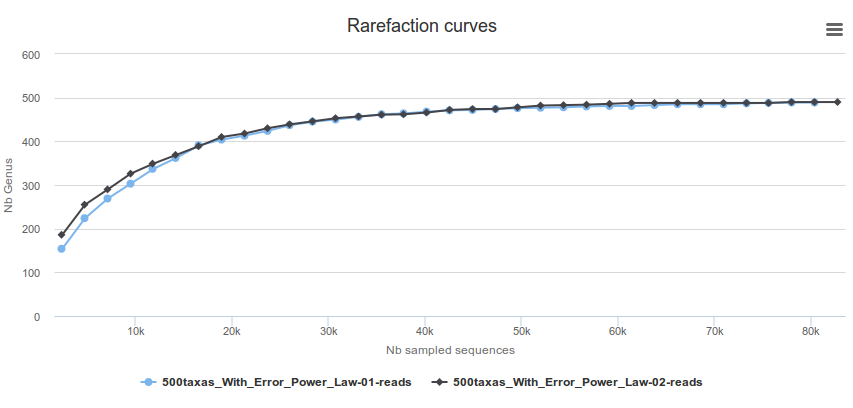
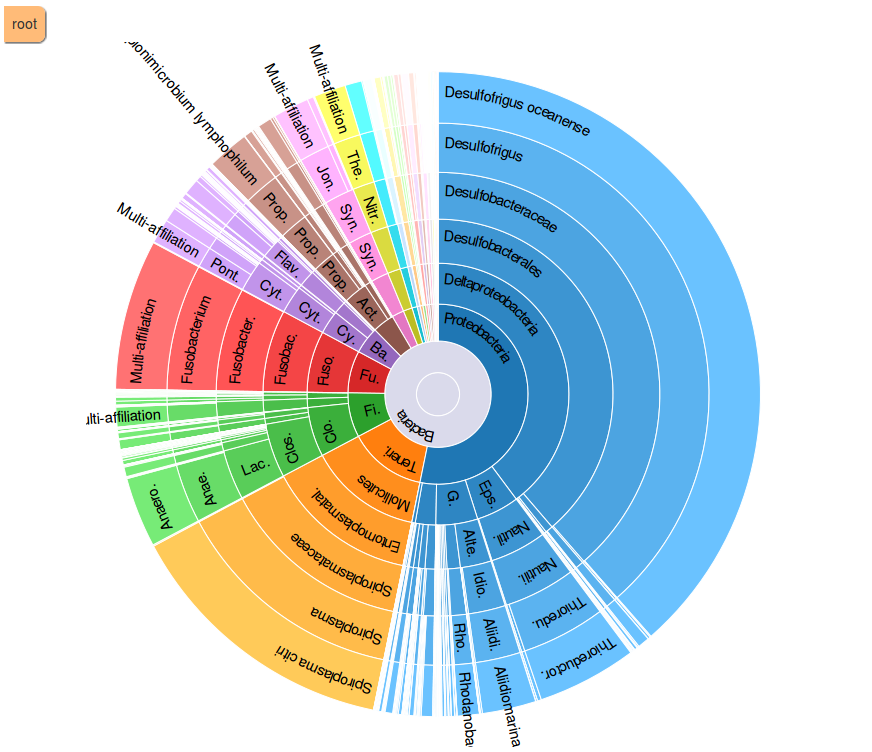
-Bootstrap distribution: display for affiliation methods with bootstrap the bootstrap on each taxonomic rank
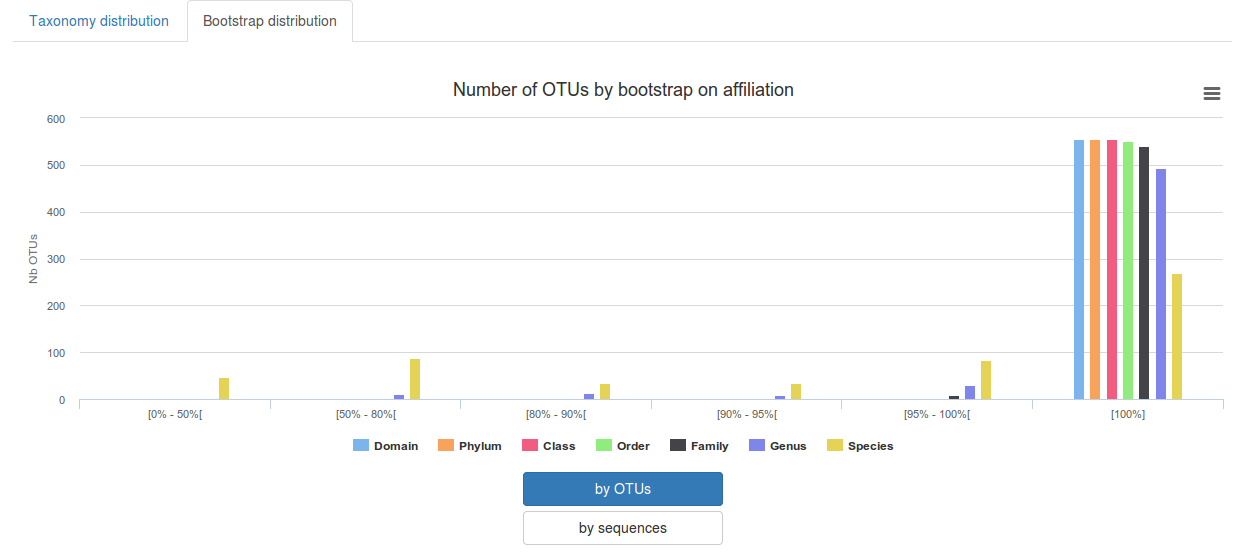
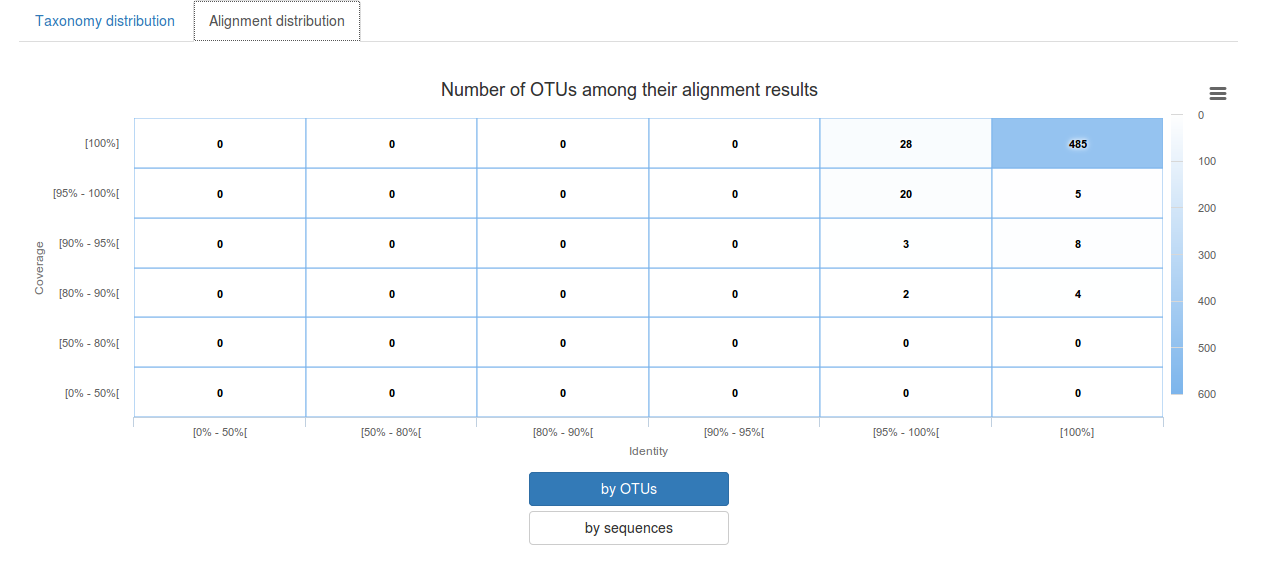
-Alignment distribution: display for affiliation methods with alignment the distribution of identity/covrage
Contact
Contacts: frogs@inra.fr
Repository: https://github.com/geraldinepascal/FROGS
Please cite the FROGS Publication: Escudie F., Auer L., Bernard M., Cauquil L., Vidal K., Maman S., Mariadassou M., Hernadez-Raquet G., Pascal G., 2015. FROGS: Find Rapidly OTU with Galaxy Solution. In: The environmental genomic Conference, Montpellier, France, http://bioinfo.genotoul.fr/fileadmin/user_upload/FROGS_2015_GE_Montpellier_poster.pdf
Depending on the help provided you can cite us in acknowledgements, references or both.