Arriba Draw Fusions
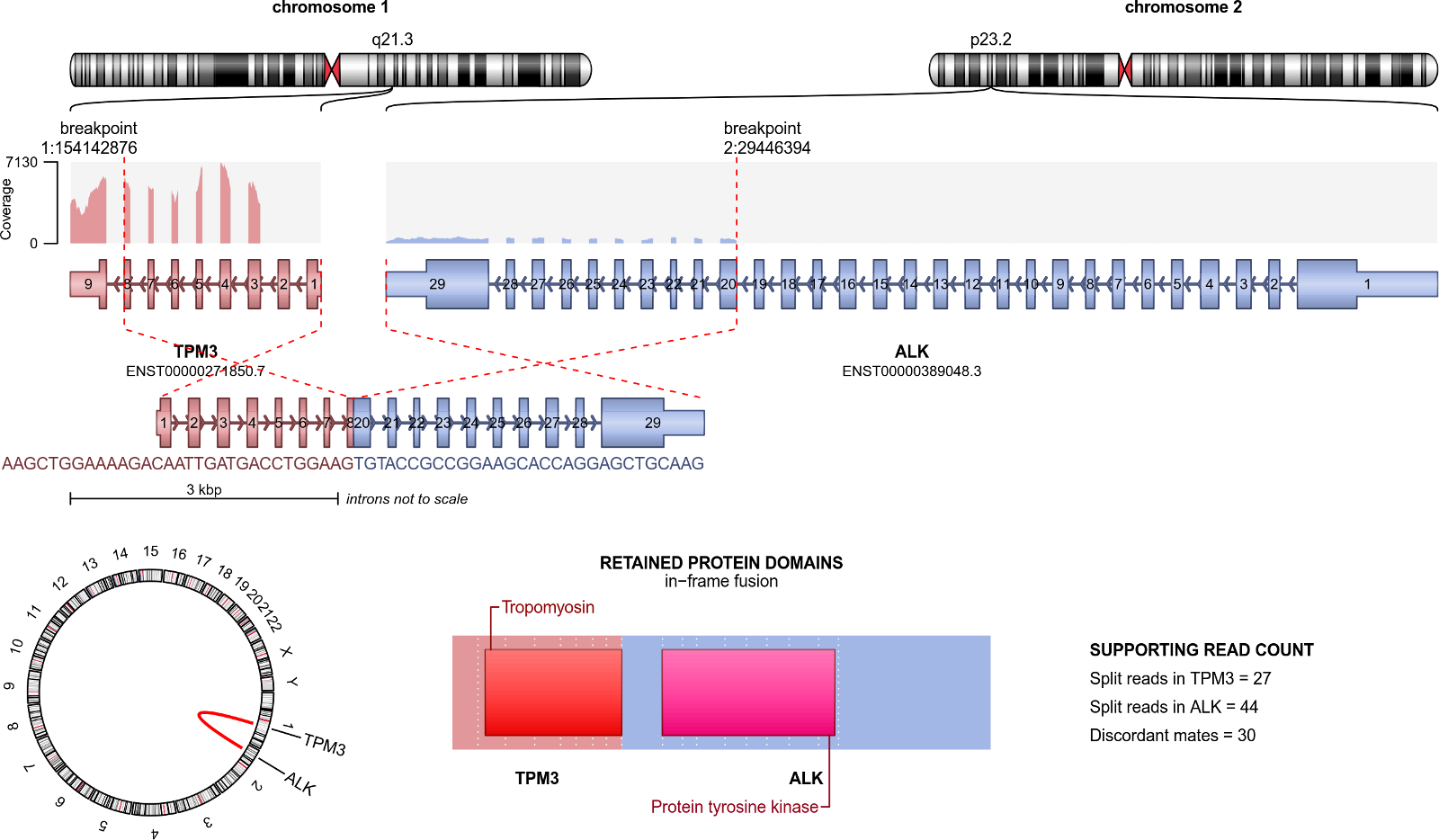
Arriba_Draw_Fusions (draw_fusions.R) renders publication-quality visualizations of the transcripts involved in predicted fusions. It generates a PDF file with one page for each predicted fusion. Each page depicts the fusion partners, their orientation, the retained exons in the fusion transcript, statistics about the number of supporting reads, and - if the column fusion_transcript has a value - an excerpt of the sequence around the breakpoint.
INPUTS
See: https://arriba.readthedocs.io/en/latest/command-line-options/#draw_fusionsr
Fusions
File containing fusion predictions from Arriba (fusions.tsv) or STAR-Fusion (star-fusion.fusion_predictions.tsv or star-fusion.fusion_predictions.abridged.coding_effect.tsv).
Annotation
Gene annotation in GTF format that was used by the STAR aligner.
Alignments
BAM file containing normal alignments from STAR.
Annotation
The gene annotation (parameter -g) is used for multiple purposes:
Assembly (Optional)
Only required when alignments are not sorted bam format. The genonme assembly will be used by samtools to produce a sorted bam file.
Protein domains (Optional)
GFF3 file containing the genomic coordinates of protein domains. Distributions of Arriba offer protein domain annotations for all supported assemblies in the database directory. When this file is given, a plot is generated, which shows the protein domains retained in the fusion transcript.
Cytobands (Optional)
Coordinates of the Giemsa staining bands. This information is used to draw ideograms. If the argument is omitted, then no ideograms are rendered. The file must have the following columns: contig, start, end, name, giemsa. Recognized values for the Giemsa staining intensity are: gneg, gpos followed by a percentage, acen, stalk. Cytobands forahuman and mouse reference can be retrieved from the Arriba distribution with the Arriba Get Filters tool.
OPTIONS
See: https://arriba.readthedocs.io/en/latest/command-line-options/#draw_fusionsr
OUTPUTS
See: https://arriba.readthedocs.io/en/latest/visualization/
fusions.pdf
A PDF file with one page for each predicted fusion. Each page depicts the fusion partners, their orientation, the retained exons in the fusion transcript, statistics about the number of supporting reads, and if the column fusion_transcript has a value an excerpt of the sequence around the breakpoint.