What it does
Interproscan is a batch tool to query the Interpro database. It provides annotations based on multiple searches of profile and other functional databases.
Input
Required is a FASTA file containing protein or nucleotide sequences.
Output
In this version of InterProScan, you can retrieve output in any of the following five formats:
Tab-separated values format (TSV)
Basic tab delimited format.
Example Output
P51587 14086411a2cdf1c4cba63020e1622579 3418 Pfam PF09103 BRCA2, oligonucleotide/oligosaccharide-binding, domain 1 2670 2799 7.9E-43 T 15-03-2013 P51587 14086411a2cdf1c4cba63020e1622579 3418 ProSiteProfiles PS50138 BRCA2 repeat profile. 1002 1036 0.0 T 18-03-2013 IPR002093 BRCA2 repeat GO:0005515|GO:0006302 P51587 14086411a2cdf1c4cba63020e1622579 3418 Gene3D G3DSA:2.40.50.140 2966 3051 3.1E-52 T 15-03-2013 ...
The TSV format presents the match data in columns as follows:
- Protein Accession (e.g. P51587)
- Sequence MD5 digest (e.g. 14086411a2cdf1c4cba63020e1622579)
- Sequence Length (e.g. 3418)
- Analysis (e.g. Pfam / PRINTS / Gene3D)
- Signature Accession (e.g. PF09103 / G3DSA:2.40.50.140)
- Signature Description (e.g. BRCA2 repeat profile)
- Start location
- Stop location
- Score - is the e-value of the match reported by member database method (e.g. 3.1E-52)
- Status - is the status of the match (T: true)
- Date - is the date of the run
- (InterProScan annotations - accession (e.g. IPR002093) - optional column; only displayed if -iprscan option is switched on)
- (InterProScan annotations - description (e.g. BRCA2 repeat) - optional column; only displayed if -iprscan option is switched on)
- (GO annotations (e.g. GO:0005515) - optional column; only displayed if --goterms option is switched on)
- (Pathways annotations (e.g. REACT_71) - optional column; only displayed if --pathways option is switched on)
Extensible Markup Language (XML)
XML representation of the matches - this is the richest form of the data. The XML Schema Definition (XSD) is available [http://www.ebi.ac.uk/interpro/resources/schemas/interproscan5 here].
Example Output
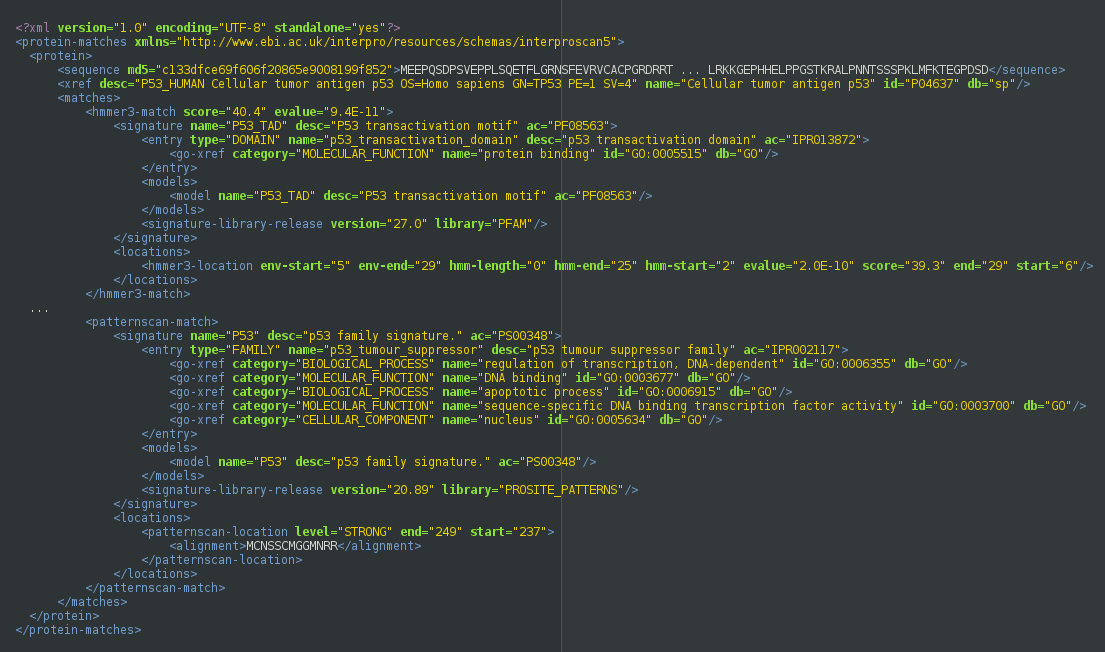
Generic Feature Format Version 3 (GFF3)
The GFF3 format is a flat tab-delimited file, which is much richer then the TSV output format. It allows you to trace back from matches to predicted proteins and to nucleic acid sequences. It also contains a FASTA format representation of the predicted protein sequences and their matches. You will find a documentation of all the columns and attributes used on [http://www.sequenceontology.org/gff3.shtml].
Example Output
AACH01000027 provided_by_user nucleic_acid 1 1347 . + . Name=AACH01000027;md5=b2a7416cb92565c004becb7510f46840;ID=AACH01000027 AACH01000027 getorf ORF 1 1347 . + . Name=AACH01000027.2_21;Target=pep_AACH01000027_1_1347 1 449;md5=b2a7416cb92565c004becb7510f46840;ID=orf_AACH01000027_1_1347 AACH01000027 getorf polypeptide 1 449 . + . md5=fd0743a673ac69fb6e5c67a48f264dd5;ID=pep_AACH01000027_1_1347 AACH01000027 Pfam protein_match 84 314 1.2E-45 + . Name=PF00696;signature_desc=Amino acid kinase family;Target=null 84 314;status=T;ID=match$8_84_314;Ontology_term="GO:0008652";date=15-04-2013;Dbxref="InterPro:IPR001048","Reactome:REACT_13" ... >pep_AACH01000027_1_1347 LVLLAAFDCIDDTKLVKQIIISEIINSLPNIVNDKYGRKVLLYLLSPRDPAHTVREIIEV LQKGDGNAHSKKDTEIRRREMKYKRIVFKVGTSSLTNEDGSLSRSKVKDITQQLAMLHEA GHELILVSSGAIAAGFGALGFKKRPTKIADKQASAAVGQGLLLEEYTTNLLLRQIVSAQI LLTQDDFVDKRRYKNAHQALSVLLNRGAIPIINENDSVVIDELKVGDNDTLSAQVAAMVQ ADLLVFLTDVDGLYTGNPNSDPRAKRLERIETINREIIDMAGGAGSSNGTGGMLTKIKAA TIATESGVPVYICSSLKSDSMIEAAEETEDGSYFVAQEKGLRTQKQWLAFYAQSQGSIWV DKGAAEALSQYGKSLLLSGIVEAEGVFSYGDIVTVFDKESGKSLGKGRVQFGASALEDML RSQKAKGVLIYRDDWISITPEIQLLFTEF ... >match$8_84_314 KRIVFKVGTSSLTNEDGSLSRSKVKDITQQLAMLHEAGHELILVSSGAIAAGFGALGFKK RPTKIADKQASAAVGQGLLLEEYTTNLLLRQIVSAQILLTQDDFVDKRRYKNAHQALSVL LNRGAIPIINENDSVVIDELKVGDNDTLSAQVAAMVQADLLVFLTDVDGLYTGNPNSDPR AKRLERIETINREIIDMAGGAGSSNGTGGMLTKIKAATIATESGVPVYICS
Scalable Vector Graphics (SVG) and HyperText Markup Language (HTML)
InterProScan 5 outputs a single HTML/SVG file for each protein sequence analysed.
Example Output
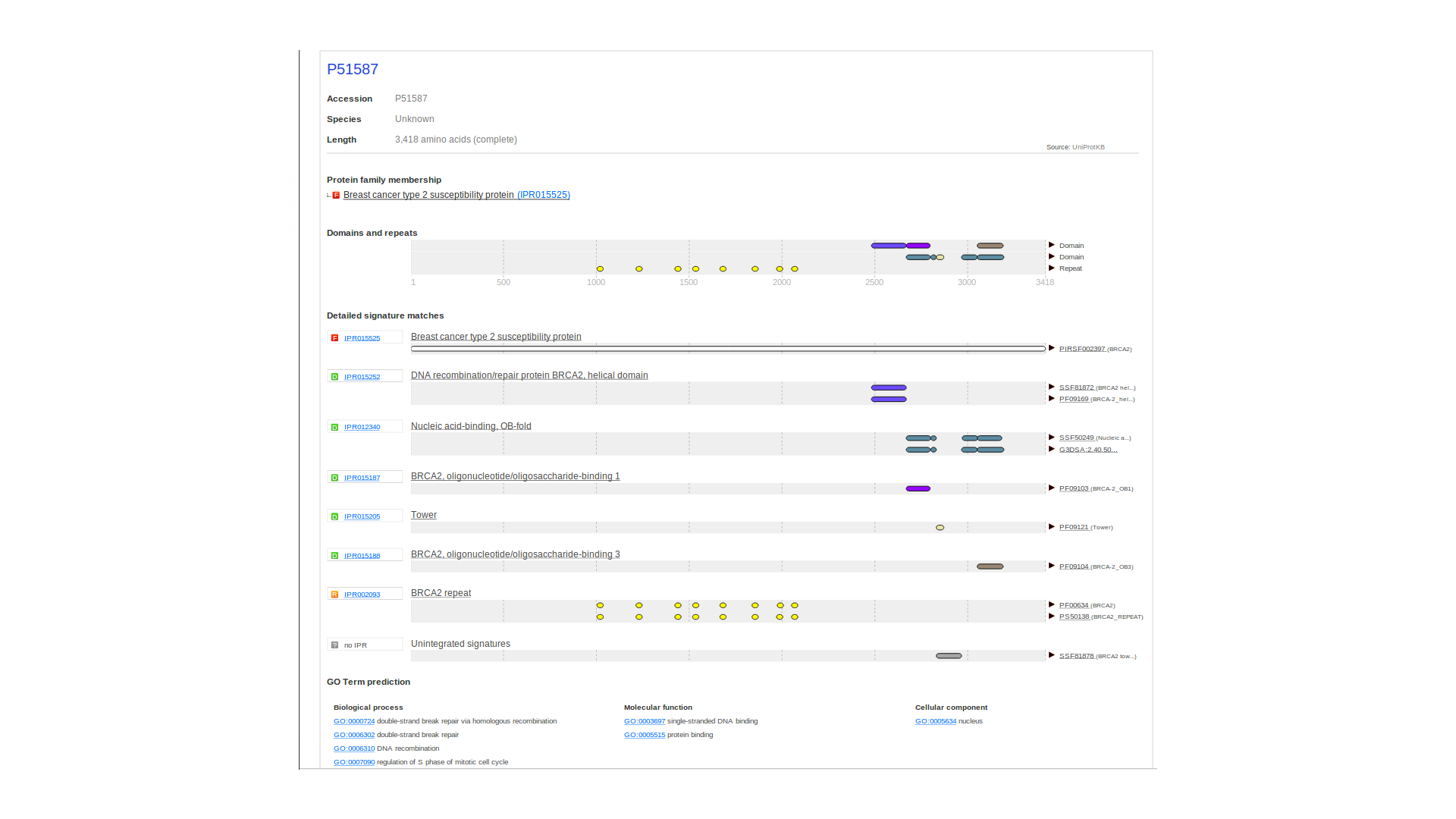
References
If you use this Galaxy tool in work leading to a scientific publication please cite the following papers:
Peter J.A. Cock, Björn A. Grüning, Konrad Paszkiewicz and Leighton Pritchard (2013). Galaxy tools and workflows for sequence analysis with applications in molecular plant pathology. PeerJ 1:e167 http://dx.doi.org/10.7717/peerj.167
Zdobnov EM, Apweiler R (2001) InterProScan an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17, 847-848. http://dx.doi.org/10.1093/bioinformatics/17.9.847
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Research 33 (Web Server issue), W116-W120. http://dx.doi.org/10.1093/nar/gki442
Hunter S, Apweiler R, Attwood TK, Bairoch A, Bateman A, Binns D, Bork P, Das U, Daugherty L, Duquenne L, Finn RD, Gough J, Haft D, Hulo N, Kahn D, Kelly E, Laugraud A, Letunic I, Lonsdale D, Lopez R, Madera M, Maslen J, McAnulla C, McDowall J, Mistry J, Mitchell A, Mulder N, Natale D, Orengo C, Quinn AF, Selengut JD, Sigrist CJ, Thimma M, Thomas PD, Valentin F, Wilson D, Wu CH, Yeats C. (2009) InterPro: the integrative protein signature database. Nucleic Acids Research 37 (Database Issue), D224-228. http://dx.doi.org/10.1093/nar/gkn785
This wrapper is available to install into other Galaxy Instances via the Galaxy Tool Shed at http://toolshed.g2.bx.psu.edu/view/bgruening/interproscan5
Galaxy Wrapper Author:
* Bjoern Gruening, University of Freiburg * Konrad Paszkiewicz, University of Exeter